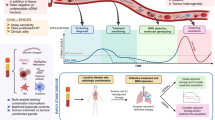
Abstract
Upfront tumour genotyping is now considered an essential step in guiding treatment decision-making in the management of patients with advanced-stage non-small-cell lung cancer (NSCLC) in light of the ever-expanding toolbox of targeted therapies and immune-checkpoint inhibitors. However, genotyping of tumour biopsy samples is not feasible for all patients and, therefore, genomic analysis of circulating tumour DNA (ctDNA) has emerged as a compelling non-invasive option. Current guidelines universally recommend genotyping and support the use of ctDNA testing in certain settings, although they often omit the detail necessary for integrating these tests into clinical care on an individual basis. In this Perspective, we describe the rationale, promise and challenges associated with ctDNA-based NSCLC genotyping and suggest a framework for the implementation of these assays into routine clinical practice. We also offer considerations for the interpretation of ctDNA genotyping results, which, particularly when using next-generation sequencing panels, can be nuanced. Through the addition of this new approach to clinical practice, we propose that oncologists might finally be able to utilize effective genotyping in nearly all patients with advanced-stage NSCLC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Carlisle, J. W. & Ramalingam, S. S. A banner year for immunotherapy and targeted therapy. Nat. Rev. Clin. Oncol. 16, 79–80 (2019).
Lisberg, A. et al. A phase II study of pembrolizumab in EGFR-mutant, PD-L1+, tyrosine kinase inhibitor naïve patients with advanced NSCLC. J. Thorac. Oncol. 13, 1138–1145 (2018).
Mazieres, J. et al. Immune checkpoint inhibitors for patients with advanced lung cancer and oncogenic driver alterations: results from the IMMUNOTARGET registry. Ann. Oncol. 30, 1321–1328 (2019).
Brown, H. et al. Programmed cell death ligand 1 expression in untreated EGFR mutated advanced NSCLC and response to osimertinib versus comparator in FLAURA. J. Thorac. Oncol. 15, 138–143 (2020).
Mok, T. S. K. et al. Pembrolizumab versus chemotherapy for previously untreated, PD-L1-expressing, locally advanced or metastatic non-small-cell lung cancer (KEYNOTE-042): a randomised, open-label, controlled, phase 3 trial. Lancet 393, 1819–1830 (2019).
Reck, M. et al. Pembrolizumab versus chemotherapy for PD-L1–positive non–small-cell lung cancer. N. Engl. J. Med. 375, 1823–1833 (2016).
Lindeman, N. I. et al. Updated molecular testing guideline for the selection of lung cancer patients for treatment with targeted tyrosine kinase inhibitors: guideline from the College of American Pathologists, the International Association for the Study of Lung Cancer, and the Association for Molecular Pathology. J. Thorac. Oncol. 13, 323–358 (2018).
Ettinger, D. S. et al. NCCN guidelines insights: non–small cell lung cancer, version 1.2020. J. Natl Compr. Canc. Netw. 17, 1464 (2019).
Kalemkerian, G. P. et al. Molecular testing guideline for the selection of patients with lung cancer for treatment with targeted tyrosine kinase inhibitors: American Society of Clinical Oncology Endorsement of the College of American Pathologists/International Association for the Study of Lung Cancer/Association for Molecular Pathology Clinical Practice guideline update. J. Clin. Oncol. 36, 911–919 (2018).
Pennell, N. A. et al. Economic impact of next-generation sequencing versus single-gene testing to detect genomic alterations in metastatic non–small-cell lung cancer using a decision analytic model. JCO Precis. Oncol. 3, 1–9 (2019).
Sholl, L. M. et al. Institutional implementation of clinical tumor profiling on an unselected cancer population. JCI Insight 1, e87062 (2016).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Tan, A. C. et al. Utility of incorporating next-generation sequencing (NGS) in an Asian non-small cell lung cancer (NSCLC) population: Incremental yield of actionable alterations and cost-effectiveness analysis. Lung Cancer 139, 207–215 (2020).
Aggarwal, C. et al. Clinical implications of plasma-based genotyping with the delivery of personalized therapy in metastatic non–small cell lung cancer. JAMA Oncol. 5, 173–180 (2019).
Bettegowda, C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl Med. 6, 224ra224 (2014).
Fan, H. C., Blumenfeld, Y. J., Chitkara, U., Hudgins, L. & Quake, S. R. Noninvasive diagnosis of fetal aneuploidy by shotgun sequencing DNA from maternal blood. Proc. Natl Acad. Sci. USA 105, 16266–16271 (2008).
Kapeleris, J. et al. The prognostic role of circulating tumor cells (CTCs) in lung cancer. Front. Oncol. 8, 311 (2018).
Russo, A. et al. Liquid biopsy tracking of lung tumor evolutions over time. Expert. Rev. Mol. Diagn. 19, 1099–1108 (2019).
Yung, T. K. F. et al. Single-molecule detection of epidermal growth factor receptor mutations in plasma by microfluidics digital PCR in non–small cell lung cancer patients. Clin. Cancer Res. 15, 2076–2084 (2009).
Diehl, F. et al. Detection and quantification of mutations in the plasma of patients with colorectal tumors. Proc. Natl Acad. Sci. USA 102, 16368–16373 (2005).
Oxnard, G. R., Paweletz, C. P. & Sholl, L. M. Genomic analysis of plasma cell-free DNA in patients with cancer. JAMA Oncol. 3, 740–741 (2017).
Wu, Y.-L. et al. Clinical utility of a blood-based EGFR mutation test in patients receiving first-line erlotinib therapy in the ENSURE, FASTACT-2, and ASPIRATION studies. Lung Cancer 126, 1–8 (2018).
Jenkins, S. et al. Plasma ctDNA analysis for detection of the EGFR T790M mutation in patients with advanced non–small cell lung cancer. J. Thorac. Oncol. 12, 1061–1070 (2017).
Sacher, A. G. et al. Prospective validation of rapid plasma genotyping for the detection of EGFR and KRAS mutations in advanced lung cancer. JAMA Oncol. 2, 1014–1022 (2016).
Remon, J. et al. Real-world utility of an amplicon-based next-generation sequencing liquid biopsy for broad molecular profiling in patients with advanced non–small-cell lung cancer. JCO Precis. Oncol. https://doi.org/10.1200/po.18.00211 (2019).
Paweletz, C. P. et al. Bias-corrected targeted next-generation sequencing for rapid, multiplexed detection of actionable alterations in cell-free DNA from advanced lung cancer patients. Clin. Cancer Res. 22, 915–922 (2016).
Schrock, A. B. et al. Hybrid capture–based genomic profiling of circulating tumor DNA from patients with advanced non–small cell lung cancer. J. Thorac. Oncol. 14, 255–264 (2019).
Müller, J. N. et al. Concordance between comprehensive cancer genome profiling in plasma and tumor specimens. J. Thorac. Oncol. 12, 1503–1511 (2017).
Cheng, J. et al. Clinical validation of a cell-free DNA gene panel. J. Mol. Diagn. 21, 632–645 (2019).
Leighl, N. B. et al. Clinical utility of comprehensive cell-free DNA analysis to identify genomic biomarkers in patients with newly diagnosed metastatic non–small cell lung cancer. Clin. Cancer Res. 25, 4691–4700 (2019).
Gadgeel, S. M. et al. Phase II/III blood first assay screening trial (BFAST) in patients (pts) with treatment-naïve NSCLC: initial results from the ALK+ cohort. Ann. Oncol. 30 (Suppl. 5), v851–v934 (2019).
Hu, Y. et al. False-positive plasma genotyping due to clonal hematopoiesis. Clin. Cancer Res. 24, 4437–4443 (2018).
Razavi, P. et al. High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat. Med. 25, 1928–1937 (2019).
Li, B. T. et al. Ultra-deep next-generation sequencing of plasma cell-free DNA in patients with advanced lung cancers: results from the Actionable Genome Consortium. Ann. Oncol. 30, 597–603 (2019).
Gray, J. E. et al. Tissue and plasma EGFR mutation analysis in the FLAURA trial: osimertinib versus comparator EGFR tyrosine kinase inhibitor as first-line treatment in patients with EGFR-mutated advanced non–small cell lung cancer. Clin. Cancer Res. 25, 6644–6652 (2019).
Passiglia, F et al. Metastatic site location influences the diagnostic accuracy of ctDNA EGFR- mutation testing in NSCLC patients: a pooled analysis. Curr. Cancer Drug Targets 18, 697–705 (2018).
Meador, C. et al. Refining the sensitivity of plasma cell-free DNA (cfDNA) genotyping by controlling for plasma tumor content. J. Clin. Oncol. 36, 9071–9071 (2018).
Supplee, J. G. et al. Sensitivity of next-generation sequencing assays detecting oncogenic fusions in plasma cell-free DNA. Lung Cancer 134, 96–99 (2019).
Dagogo-Jack, I. et al. Molecular analysis of plasma from patients with ROS1-positive NSCLC. J. Thorac. Oncol. 14, 816–824 (2019).
Oxnard, G. R. et al. Association between plasma genotyping and outcomes of treatment with osimertinib (AZD9291) in advanced non-small-cell lung cancer. J. Clin. Oncol. 34, 3375–3382 (2016).
Oxnard, G. R. et al. Assessment of resistance mechanisms and clinical implications in patients with EGFR T790M–positive lung cancer and acquired resistance to osimertinib. JAMA Oncol. 4, 1527–1534 (2018).
Siravegna, G. et al. Clonal evolution and resistance to EGFR blockade in the blood of colorectal cancer patients. Nat. Med. 21, 795–801 (2015).
Pectasides, E. et al. Genomic heterogeneity as a barrier to precision medicine in gastroesophageal adenocarcinoma. Cancer Discov. 8, 37–48 (2018).
Parikh, A. R. et al. Liquid versus tissue biopsy for detecting acquired resistance and tumor heterogeneity in gastrointestinal cancers. Nat. Med. 25, 1415–1421 (2019).
Bai, H. et al. Detection and clinical significance of intratumoral EGFR mutational heterogeneity in chinese patients with advanced non-small cell lung cancer. PLoS ONE 8, e54170 (2013).
Yatabe, Y., Matsuo, K. & Mitsudomi, T. Heterogeneous distribution of EGFR mutations is extremely rare in lung adenocarcinoma. J. Clin. Oncol. 29, 2972–2977 (2011).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Gandara, D. R. et al. Blood-based tumor mutational burden as a predictor of clinical benefit in non-small-cell lung cancer patients treated with atezolizumab. Nat. Med. 24, 1441–1448 (2018).
Rolfo, C. et al. Liquid biopsy for advanced non-small cell lung cancer (NSCLC): a statement paper from the IASLC. J. Thorac. Oncol. 13, 1248–1268 (2018).
Roche. cobas® EGFR Mutation Test v2 https://diagnostics.roche.com/us/en/products/params/cobas-egfr-mutation-test-v2.html (2020).
Sabari, J. K. et al. A prospective study of circulating tumor DNA to guide matched targeted therapy in lung cancers. J. Natl Cancer Inst. 111, 575–583 (2018).
Sorber, L. et al. Specialized blood collection tubes for liquid biopsy: improving the pre-analytical conditions. Mol. Diagn. Ther. 24, 113–124 (2020).
Siravegna, G., Marsoni, S., Siena, S. & Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol. 14, 531–548 (2017).
Travis, W. D. et al. Diagnosis of lung cancer in small biopsies and cytology: implications of the 2011 International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society Classification. Arch. Pathol. Lab. Med. 137, 668–684 (2013).
Trisolini, R. et al. Randomized trial of endobronchial ultrasound-guided transbronchial needle aspiration with and without rapid on-site evaluation for lung cancer genotyping. Chest 148, 1430–1437 (2015).
Zill, O. A. et al. The landscape of actionable genomic alterations in cell-free circulating tumor DNA from 21,807 advanced cancer patients. Clin. Cancer Res. 24, 3528–3538 (2018).
Merker, J. D. et al. Circulating tumor DNA analysis in patients with cancer: American Society of Clinical Oncology and College of American Pathologists joint review. J. Clin. Oncol. 36, 1631–1641 (2018).
Vu, P., Khagi, Y., Riviere, P., Goodman, A. & Kurzrock, R. Total number of alterations in liquid biopsies is an independent predictor of survival in patients with advanced cancers. JCO Precis. Oncol. 4, 192–201 (2020).
Hu, Y. et al. Discrimination of germline EGFR T790M mutations in plasma cell-free DNA allows study of prevalence across 31,414 cancer patients. Clin. Cancer Res. 23, 7351–7359 (2017).
Slavin, T. P. et al. Identification of incidental germline mutations in patients with advanced solid tumors who underwent cell-free circulating tumor DNA sequencing. J. Clin. Oncol. 36, 3459–3465 (2018).
Acknowledgements
G.R.O. is the Damon Runyon-Gordon Family Clinical Investigator supported by the Damon Runyon Cancer Research Foundation (CI-86-16) and is supported in part by the National Cancer Institute of the NIH (R01CA240592, P30CA093373). The authors wish to thank AstraZeneca and Guardant Health for providing timely information regarding regulatory developments, which helped to make this Perspective possible.
Author information
Authors and Affiliations
Contributions
G.R.O. researched data for this manuscript, all authors made a substantial contribution to discussions of content, and all authors wrote the manuscript and edited and/or reviewed the manuscript prior to submission.
Corresponding author
Ethics declarations
Competing interests
C.A. has received consultancy fees or honoraria from AstraZeneca, BMS, Celgene, Lilly, Merck, and Roche-Genentech and institutional research funding from Macrogenics, Merck, Novartis, and Xencor. C.D.R. has received honoraria from AstraZeneca and MSD, consultancy fees from Archer, EMD, Inivata, Mylan, and Serono and has acted as an adviser (non-compensated) of Oncompass. G.R.O. became a full-time employee of Foundation Medicine after submission of this manuscript and has received consultancy fees or honoraria from Abbvie, AstraZeneca, Blueprint, Dropworks, Foundation Medicine, GRAIL, Guardant Health, Illumina, Inivata, Janssen, Loxo Oncology, Sysmex, and Takeda and institutional research funding from GRAIL. J.E.G. has received consultancy fees from AstraZeneca, BMS, EMD Serono/Merck KGaA, Inivata, Merck, and Novartis and institutional research funding from Array, AstraZeneca, BMS, Boehringer–Ingelheim, Merck and Roche-Genentech. L.M.S. has received consultancy fees or honoraria from AstraZeneca, EMD Serono, Foghorn Therapeutics, and Loxo Oncology and institutional research funding from Roche Genentech. D.R.G. has received consultancy fees from AstraZeneca, Celgene, FujiFilm, Guardant Health, Inivata, IO Biotech, Lilly, Merck, Oncocyte, Roche-Genentech, and Samsung Bioepis and institutional research funding from Amgen, Astex, Merck, and Roche-Genentech.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Aggarwal, C., Rolfo, C.D., Oxnard, G.R. et al. Strategies for the successful implementation of plasma-based NSCLC genotyping in clinical practice. Nat Rev Clin Oncol 18, 56–62 (2021). https://doi.org/10.1038/s41571-020-0423-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41571-020-0423-x
This article is cited by
-
Exploring non-invasive precision treatment in non-small cell lung cancer patients through deep learning radiomics across imaging features and molecular phenotypes
Biomarker Research (2024)
-
Plasma cell-free DNA as a sensitive biomarker for multi-cancer detection and immunotherapy outcomes prediction
Journal of Cancer Research and Clinical Oncology (2024)
-
The Impact of Prior Single-Gene Testing on Comprehensive Genomic Profiling Results for Patients with Non-Small Cell Lung Cancer
Oncology and Therapy (2024)
-
Monitoring of T790M in plasma ctDNA of advanced EGFR-mutant NSCLC patients on first- or second-generation tyrosine kinase inhibitors
BMC Cancer (2023)
-
Plasma ctDNA increases tissue NGS-based detection of therapeutically targetable mutations in lung cancers
BMC Cancer (2023)