Abstract
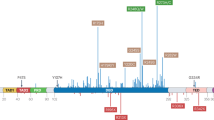
The tumor suppressor protein p53, a transcription product of the anti-oncogene TP53, is a critical factor in preventing cellular cancerization and killing cancer cells by inducing apoptosis. As a result, p53 is often referred to as the “guardian of the genome.” Almost half of cancers possess genetic mutations in the TP53 gene, and most of these mutations result in the malfunction of p53, which promotes aggregation. In some cases, the product of the TP53 mutant allele shows higher aggregation propensity; the mutant co-aggregates with the normal (functional) p53 protein, thus losing cellular activity of the p53 guardian. Cancer might also progress because of the proteolytic degradation of p53 by activated E3 ubiquitination enzymes, MDM2 and MDM4. The inhibition of the specific interaction between MDM2 (MDM4) and p53 also results in increased p53 activity in cancer cells. Although the molecular targets of the drugs are different, two drug discovery strategies with a common goal, “rescuing p53 protein,” have recently emerged. To conduct this approach, various biophysical methods of protein characterization were employed. In this review, we focus on these two independent strategies based on the unique biophysical features of the p53 protein.


Similar content being viewed by others
Availability of data and material
Not applicable.
Code availability
Not applicable.
Abbreviations
- Aβ:
-
Amyloid-β peptide
- DBD:
-
DNA-binding domain
- HDM2:
-
Human double minutes 2
- HTS:
-
High-throughput screening
- IARC:
-
International Agency for Research on Cancer
- IDR:
-
Intrinsically disordered regions
- LLPS:
-
Liquid–liquid phase separation
- MDM2:
-
Murine double minutes 2
- MQ:
-
2-Methylene-3-quinuclidinone
- OD:
-
Oligomer domain
- PRIMA-1:
-
p53 reactivation with induction of massive apoptosis-1
- REG:
-
C-terminal regulatory domain
- TAD:
-
Transactivation domain
- TCGA:
-
The Cancer Genome Atlas
- TD:
-
Tetramerization domain
References
Ano Bom APD, Rangel LP, Costa DCF et al (2012) Mutant p53 aggregates into prion-like amyloid oligomers and fibrils: implications for cancer. J Biol Chem 287:28152–28162. https://doi.org/10.1074/jbc.M112.340638
Bauer MR, Jones RN, Tareque RK et al (2019) A structure-guided molecular chaperone approach for restoring the transcriptional activity of the p53 cancer mutant Y220C. Future Med Chem 11:2491–2504. https://doi.org/10.4155/fmc-2019-0181
Baugh EH, Ke H, Levine AJ et al (2018) Why are there hotspot mutations in the TP53 gene in human cancers? Cell Death Differ 25:154–160. https://doi.org/10.1038/cdd.2017.180
Billant O, Friocourt G, Roux P, Voisset C (2021) p53, a victim of the prion fashion. Cancers (basel) 13:1–15. https://doi.org/10.3390/cancers13020269
Boeckler F, Joerger A, Jaggi G et al (2008) Targeted rescue of a destabilized mutant of p53 by an in silico screened drug. Proc Natl Acad Sci U S A 105:10360–10365. https://doi.org/10.1073/pnas.0805326105
Bouaoun L, Sonkin D, Ardin M et al (2016) TP53 variations in human cancers: new lessons from the IARC TP53 database and genomics data. Hum Mutat 37:865–876. https://doi.org/10.1002/humu.23035
Bullock AN, Fersht AR (2001) Rescuing the function of mutant p53. Nat Rev Cancer 1:68–76. https://doi.org/10.1038/35094077
Bullock AN, Henckel J, Fersht AR (2000) Quantitative analysis of residual folding and DNA binding in mutant p53 core domain: definition of mutant states for rescue in cancer therapy. Oncogene 19:1245–1256. https://doi.org/10.1038/sj.onc.1203434
Bykov VJN, Issaeva N, Shilov A et al (2002) Restoration of the tumor suppressor function to mutant p53 by a low-molecular-weight compound. Nat Med 8:282–288. https://doi.org/10.1038/nm0302-282
da Costa DCF, Campos NPC, Santos RA, et al (2018) Resveratrol prevents p53 aggregation in vitro and in breast cancer cells. Oncotarget 9:29112–29122. https://doi.org/10.18632/oncotarget.25631
Davydov IV, Woods D, Safiran YJ et al (2004) Assay for ubiquitin ligase activity: high-throughput screen for inhibitors of HDM2. J Biomol Screen 9:695–703. https://doi.org/10.1177/1087057104267956
Ding Q, Zhang Z, Liu JJ et al (2013) Discovery of RG7388, a potent and selective p53-MDM2 inhibitor in clinical development. J Med Chem 56:5979–5983. https://doi.org/10.1021/jm400487c
Dyson HJ, Wright PE (2005) Intrinsically unstructured proteins and their functions. Nat Rev Mol Cell Biol 6:197–208. https://doi.org/10.1038/nrm1589
Feng Y, Wang, Yang XPSG et al (2009) Resveratrol inhibits beta-amyloid oligomeric cytotoxicity but does not prevent oligomer formation. Neurotoxicology 30:986–995. https://doi.org/10.1016/j.neuro.2009.08.013
Fukuchi S, Hosoda K, Homma K et al (2011) Binary classification of protein molecules into intrinsically disordered and ordered segments. BMC Struct Biol 11. https://doi.org/10.1186/1472-6807-11-29
Galea C, Bowman P, Kriwacki RW (2005) Disruption of an intermonomer salt bridge in the p53 tetramerization domain results in an increased propensity to form amyloid fibrils. Protein Sci 14:2993–3003. https://doi.org/10.1110/ps.051622005
Garg A, Hazra JP, Sannigrahi MK et al (2020) Variable mutations at the p53–R273 oncogenic hotspot position leads to altered properties. Biophys J 118:720–728. https://doi.org/10.1016/j.bpj.2019.12.015
Ghosh S, Salot S, Sengupta S et al (2017) p53 amyloid formation leading to its loss of function: implications in cancer pathogenesis. Cell Death Differ 24:1784–1798. https://doi.org/10.1038/cdd.2017.105
Ishimaru D, Andrade LR, Teixeira LSP et al (2003a) Fibrillar aggregates of the tumor suppressor p53 core domain. Biochemistry 42:9022–9027. https://doi.org/10.1021/bi034218k
Ishimaru D, Maia LF, Maiolino LM et al (2003b) Conversion of wild-type p53 core domain into a conformation that mimics a hot-spot mutant. J Mol Biol 333:443–451. https://doi.org/10.1016/j.jmb.2003.08.026
Itoh Y, Murata A, Sakamoto S et al (2016) Activation of p53 facilitates the target search in DNA by enhancing the target recognition probability. J Mol Biol 428:2916–2930. https://doi.org/10.1016/j.jmb.2016.06.001
Iwahashi N, Ikezaki M, Nishikawa T et al (2020) Sulfated glycosaminoglycans mediate prion-like behavior of p53 aggregates. Proc Natl Acad Sci U S A 117:33225–33234. https://doi.org/10.1073/PNAS.2009931117
Jin Z, Shen J, He J, Hu C (2015) Combination therapy with p53-MDM2 binding inhibitors for malignancies. Med Chem Res 24:1369–1379. https://doi.org/10.1007/s00044-014-1089-7
Joerger AC, Fersht AR (2016) The p53 pathway: origins, inactivation in cancer, and emerging therapeutic approaches. Annu Rev Biochem 85:375–404. https://doi.org/10.1146/annurev-biochem-060815-014710
Kamagata K, Kanbayashi S, Honda M et al (2020) Liquid-like droplet formation by tumor suppressor p53 induced by multivalent electrostatic interactions between two disordered domains. Sci Rep 10:1–12. https://doi.org/10.1038/s41598-020-57521-w
Kang MH, Reynolds CP, Kolb EA et al (2016) Initial testing (stage 1) of MK-8242—a novel MDM2 inhibitor—by the pediatric preclinical testing program. Pediatr Blood Cancer 63:1744–1752. https://doi.org/10.1002/pbc.26064
Kastenhuber ER, Lowe SW (2017) Putting p53 in context. Cell 170:1062–1078. https://doi.org/10.1016/j.cell.2017.08.028
Lane DP (1992) p53, guardian of the genome. Nature 358:15–16. https://doi.org/10.1038/358015a0
Lasagna-Reeves CA, Clos AL, Castillo-Carranza D et al (2013) Dual role of p53 amyloid formation in cancer; loss of function and gain of toxicity. Biochem Biophys Res Commun 430:963–968. https://doi.org/10.1016/j.bbrc.2012.11.130
Lemos C, Schulze L, Weiske J, et al (2020) Identification of small molecules that modulate mutant p53 condensation. iScience 23. https://doi.org/10.1016/j.isci.2020.101517
Levine AJ, Oren M (2009) The first 30 years of p53: growing ever more complex. Nat Rev Cancer 9:749–758. https://doi.org/10.1038/nrc2723
Lu Y, Nikolovska-Coleska Z, Fang X et al (2006) Discovery of a nanomolar inhibitor of the human murine double minute 2 (MDM2)-p53 interaction through an integrated, virtual database screening strategy. J Med Chem 49:3759–3762. https://doi.org/10.1021/jm060023+
Marambaud P, Zhao H, Davies P (2005) Resveratrol promotes clearance of Alzheimer’s disease amyloid-β peptides. J Biol Chem 280:37377–37382. https://doi.org/10.1074/jbc.M508246200
Morimoto J, Hosono Y, Sando S (2018) Isolation of a peptide containing D-amino acid residues that inhibits the α-helix-mediated p53–MDM2 interaction from a one-bead one-compound library. Bioorganic Med Chem Lett 28:231–234. https://doi.org/10.1016/j.bmcl.2018.01.001
Nojiri T, Fukui D, Werneburg I et al (2021) Embryonic staging of bats with special reference to Vespertilio sinensis and its cochlear development. Dev Dyn 250:1140–1159. https://doi.org/10.1002/dvdy.325
Olivier M, Eeles R, Hollstein M et al (2002) The IARC TP53 database: new online mutation analysis and recommendations to users. Hum Mutat 19:607–614. https://doi.org/10.1002/humu.10081
Palanikumar L, Karpauskaite L, Al-sayegh M, et al (2021) Protein mimetic amyloid inhibitor potently abrogates cancer-associated mutant p53 aggregation and restores tumor suppressor function. Nat Commun 1–24.https://doi.org/10.1038/s41467-021-23985-1
Patel A, Lee HO, Jawerth L et al (2015) A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162:1066–1077. https://doi.org/10.1016/j.cell.2015.07.047
Pedrote MM, Motta MF, Ferretti GDS, et al (2020) Oncogenic gain of function in glioblastoma is linked to mutant p53 amyloid oligomers. iScience 23:100820. https://doi.org/10.1016/j.isci.2020.100820
Petitjean A, Mathe E, Kato S et al (2007) Impact of mutant p53 functional properties on TP53 mutation patterns and tumor phenotype: lessons from recent developments in the IARC TP53 database. Hum Mutat 28:62–629. https://doi.org/10.1002/humu.20495
Petronilho EC, Pedrote MM, Marques MA et al (2021) Phase separation of p53 precedes aggregation and is affected by oncogenic mutations and ligands. Chem Sci 12:7334–7349. https://doi.org/10.1039/d1sc01739j
Rangel LP, Ferretti GDS, Costa CL et al (2019) p53 reactivation with induction of massive apoptosis-1 (PRIMA-1) inhibits amyloid aggregation of mutant p53 in cancer cells. J Biol Chem 294:3670–3682. https://doi.org/10.1074/jbc.RA118.004671
Russo C, Osterburg C, Sirico A et al (2018) Protein aggregation of the p63 transcription factor underlies severe skin fragility in AEC syndrome. Proc Natl Acad Sci U S A 115:E906–E915. https://doi.org/10.1073/pnas.1713773115
Safari MS, Wang Z, Tailor K, et al (2019) Anomalous dense liquid condensates host the nucleation of tumor suppressor p53 fibrils. iScience 12:342–355. https://doi.org/10.1016/j.isci.2019.01.027
Schneider A, Kallen J, Ottl J et al (2021) Discovery, X-ray structure and CPP-conjugation enabled uptake of p53/MDM2 macrocyclic peptide inhibitors. RSC Chem Biol. https://doi.org/10.1039/d1cb00056j
Shin Y, Brangwynne CP (2017) Liquid phase condensation in cell physiology and disease. Science 357. https://doi.org/10.1126/science.aaf4382
Silva JL, Gallo CVDM, Costa DCF, Rangel LP (2014) Prion-like aggregation of mutant p53 in cancer. Trends Biochem Sci 39:260–267. https://doi.org/10.1016/j.tibs.2014.04.001
Soragni A, Janzen DM, Johnson LM et al (2016) A designed inhibitor of p53 aggregation rescues p53 tumor suppression in ovarian carcinomas. Cancer Cell 29:90–103. https://doi.org/10.1016/j.ccell.2015.12.002
Su A, Tabata Y, Aoki K et al (2021) Elaboration of non-naturally occurring helical tripeptides as p53-MDM2/MDMX interaction inhibitors. Chem Pharm Bull 69:681–692. https://doi.org/10.1248/cpb.c21-00238
Su A, Wang S, Sada A et al (2019) Non-naturally occurring helical molecules can interfere with p53–MDM2 and p53–MDMX protein–protein interactions. Chem Pharm Bull 67:1139–1143. https://doi.org/10.1248/cpb.c19-00501
Vassilev LT, Vu BT, Graves B, et al (2004) In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303:844–848. https://doi.org/10.1126/science.1092472
Wang GZ, Fersht AR (2017) Multisite aggregation of p53 and implications for drug rescue. Proc Natl Acad Sci U S A 114:E2634–E2643. https://doi.org/10.1073/pnas.1700308114
Wang GZ, Fersht AR (2015) Propagation of aggregated p53: cross-reaction and coaggregation vs. seeding. Proc Natl Acad Sci U S A 112:2443–2448. https://doi.org/10.1073/pnas.1500262112
Wang Z, Jensen MA, Zenklusen JC (2016) A practical guide to The Cancer Genome Atlas (TCGA). Methods Mol Biol 1418:111–141. https://doi.org/10.1007/978-1-4939-3578-9_6
Wilcken R, Wang GZ, Boeckler FM, Fersht AR (2012) Kinetic mechanism of p53 oncogenic mutant aggregation and its inhibition. Proc Natl Acad Sci U S A 109:13584–13589. https://doi.org/10.1073/pnas.1211550109
Wong KBO, DeDecker BS, Freund SMV et al (1999) Hot-spot mutants of p53 core domain evince characteristic local structural changes. Proc Natl Acad Sci U S A 96:8438–8442. https://doi.org/10.1073/pnas.96.15.8438
Xu J, Reumers J, Couceiro JR et al (2011) Gain of function of mutant p53 by coaggregation with multiple tumor suppressors. Nat Chem Biol 7:285–295. https://doi.org/10.1038/nchembio.546
Yang DS, Saeedi A, Davtyan A et al (2021) Mesoscopic protein-rich clusters host the nucleation of mutant p53 amyloid fibrils. Proc Natl Acad Sci U S A 118. https://doi.org/10.1073/pnas.2015618118
Zhang Y, Hu Y, Wang JL et al (2017) Proteomic identification of ERP29 as a key chemoresistant factor activated by the aggregating p53 mutant Arg282Trp. Oncogene 36:5473–5483. https://doi.org/10.1038/onc.2017.152
Zhang Y, Xu L, Chang Y, et al (2020) Therapeutic potential of ReACp53 targeting mutant p53 protein in CRPC. 23:160–171. https://doi.org/10.1038/s41391-019-0172-z.Therapeutic
Zhang Z, Ding Q, Liu JJ et al (2014) Discovery of potent and selective spiroindolinone MDM2 inhibitor, RO8994, for cancer therapy. Bioorganic Med Chem 22:4001–4009. https://doi.org/10.1016/j.bmc.2014.05.072
Zhou G, Pantelopulos GA, Mukherjee S, Voelz VA (2017) Bridging microscopic and macroscopic mechanisms of p53-MDM2 binding with kinetic network models. Biophys J 113:785–793. https://doi.org/10.1016/j.bpj.2017.07.009
Funding
This work was supported in part by the Japan Science and Technology Agency, a step feasibility study program (AS262Z01275Q, AS242Z00566Q), the AMED Translational Research Program, strategic promotion for practical application of innovative medical technology (TR-SPRINT) under grant number 20 lm0203011-j0002, and the Aichi Cancer Research Foundation.
Author information
Authors and Affiliations
Contributions
E. H. and H. H. wrote the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
H. H. is the founder of a Nagoya University-based spinoff startup company, BeCellBar LLC. E. H. declares that they have no conflicts of interest regarding the contents of this article.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hibino, E., Hiroaki, H. Potential of rescue and reactivation of tumor suppressor p53 for cancer therapy. Biophys Rev 14, 267–275 (2022). https://doi.org/10.1007/s12551-021-00915-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12551-021-00915-5