Abstract
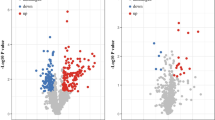
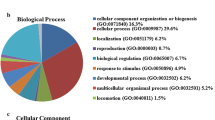
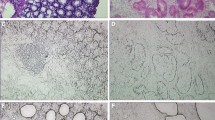
The ubiquitin–proteasome system facilitates the degradation of damaged proteins and regulators of growth and stress response. Alterations in this proteolytic system are associated with a variety of human pathologies. By restriction fragment differential display polymerase chain reaction (RFDD-PCR) and matrix-assisted laser desorption/ionization tandem time-of-flight mass spectrometry (MALDI-TOF-TOF MS) based on two-dimensional polyacrylamide gel electrophoresis (2-DE), differentially expressed genes and proteins of ubiquitin specific proteases (USPs), proteasome subuinits (PSs) and ubiquitin protein ligase E3A (UBE3A) were analyzed between breast cancer and adjacent normal tissues. Some of them were further verified as over-expression by immunohistochemical stain. Five genes of proteasome subunits (PSs), including PSMB5, PSMD1, PSMD2, PSMD8 and PSMD11, four genes of USPs, including USP9X, USP9Y, USP10 and USP25, and ubiquitin protein ligase E3A (UBE3A) were over-expressed (>3-fold) in breast cancer tissue compared to adjacent normal tissue, and over-expression (>4-fold) of proteins of PSMA1 and SMT3A were observed in breast cancer tissue. PSMD8, PSMD11 and UBE3A were further verified as over-expression by immunohistochemical stain. The action of ubiquitin–proteasome system were obviously enhanced in breast cancer, and selectively intervention in action of ubiquitin–proteasome system may be a useful method of treating human breast cancer.



Similar content being viewed by others
References
Rape M, Reddy SK, Kirschner MW (2006) The processivity of multiubiquitination by the APC determines the order of substrate degradation. Cell 124:89–103
Doucet C, Gutierrez GJ, Lindon C et al (2005) Multiple phosphorylation events control mitotic degradation of the muscle transcription factor Myf5. BMC Biochem 6:27
Asher G, Tsvetkov P, Kahana C et al (2005) A mechanism of ubiquitin-independent proteasomal degradation of the tumor suppressors p53 and p73. Genes Dev 19:316–321
Dragnev KH, Pitha-Rowe I, Ma Y et al (2004) Specific chemopreventive agents trigger proteasomal degradation of G1 cyclins: implications for combination therapy. Clin Cancer Res 10:2570–2577
Rossi S, Loda M (2003) The role of the ubiquitination-proteasome pathway in breast cancer: use of mouse models for analyzing ubiquitination processes. Breast Cancer Res 5:16–22
Zhang Q, Tian L, Mansouri A et al (2005) Inducible expression of a degradation-resistant form of p27Kip1 causes growth arrest and apoptosis in breast cancer cells. FEBS Lett 579:3932–3940
Dornan D, Bheddah S, Newton K et al (2004) COP1, the negative regulator of p53, is overexpressed in breast and ovarian adenocarcinomas. Cancer Res 64:7226–7230
Boyer L, Turchi L, Desnues B et al (2006) CNF1-induced ubiquitylation and proteasome destruction of activated RhoA is impaired in Smurf1−/− cells. Mol Biol Cell 17:2489–2497
Chen K, He M, Dong S et al (2002) Incidence, mortality and survival rates of female breast cancer in Tianjin, China. Chin J Oncol 24:573–575
Rolen U, Kobzeva V, Gasparjan N et al (2006) Activity profiling of deubiquitinating enzymes in cervical carcinoma biopsies and cell lines. Mol Carcinog 45:260–9
Ovaa H, Kessler BM, Rolen U et al (2004) Activity-based ubiquitin-specific protease (USP) profiling of virus-infected and malignant human cells. Proc Natl Acad Sci USA 101:2253–8
Dees EC, Orlowski RZ (2006) Targeting the ubiquitin–proteasome pathway in breast cancer therapy. Future Oncol 2:121–135
Ciechanover A (2005) Intracellular protein degradation: from a vague idea, through the lysosome and the ubiquitin–proteasome system, and onto human diseases and drug targeting (Nobel lecture). Angew Chem Int Ed Engl 44:5944–67
Luo Y, Zhang J, Liu Y et al (2005) Comparative proteome analysis of breast cancer and normal breast. Mol Biotechnol 29:233–244
Bennett EJ, Bence NF, Jayakumar R et al (2005) Global impairment of the ubiquitin–proteasome system by nuclear or cytoplasmic protein aggregates precedes inclusion body formation. Mol Cell 17:351–365
Ulrich HD (2002) Natural substrates of the proteasome and their recognition by the ubiquitin system. Curr Top Microbiol Immunol 268:137–174
Nijman SM, Huang TT, Dirac AM et al (2005) The deubiquitinating enzyme USP1 regulates the Fanconi anemia pathway. Mol Cell 17:331–339
Wing SS (2003) Deubiquitinating enzymes-the importance of driving in reverse along the ubiquitin–proteasome pathway. Int J Biochem Cell Biol 35:590–605
Wilkinson KD (2000) Ubiquitination and deubiquitination: targeting of proteins for degradation by the proteasome. Semin Cell Dev Biol 11:141–148
D’Andrea A, Pellman D (1998) Deubiquitinating enzymes: a new class of biological regulators. Crit Rev Biochem Mol Biol 33:337–352
Baron CA, Tepper CG, Liu SY et al (2006) Genomic and functional profiling of duplicated chromosome 15 cell lines reveal regulatory alterations in UBE3A-associated ubiquitin–proteasome pathway processes. Hum Mol Genet 15:853–869
Jiang YH, Sahoo T, Michaelis RC et al (2004) A mixed epigenetic/genetic model for oligogenic inheritance of autism with a limited role for UBE3A. Am J Med Genet A 131:1–10
Gravesen A, Warthoe P, Knochel S et al (2000) Restriction fragment differential display of pediocin-resistant Listeria monocytogenes 412 mutants shows consistent overexpression of a putative beta-glucoside-specific PTS system. Microbiology 146:1381–1389
Wrang ML, Moller F, Alsbo CW et al (2001) Changes in gene expression following induction of ischemic tolerance in rat brain: detection and verification. J Neurosci Res 65:54–58
Mei Y, Zhou HY, Xiang T et al (2005) Preliminary analysis of retinal gene expression profile of diabetic rat. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 22:563–5
Chen L, Madura K (2005) Increased proteasome activity, ubiquitin-conjugating enzymes, and eEF1A translation factor detected in breast cancer tissue. Cancer Res 65:5599–606
Nandi D, Tahiliani P, Kumar A et al (2006) The ubiquitin–proteasome system. J Biosci 31:137–55
Semple CA, RIKEN GER Group, GSL Members (2003) The comparative proteomics of ubiquitination in mouse. Genome Res 13:1389–1394
Groll M, Ditzel L, Lowe J et al (1997) Structure of 20S proteasome from yeast at 2.4 Å resolution. Nature 386:463–71
Wang H, Liu S, Lu X et al (2003) Gene expression profile changes in NB4 cells induced by realgar. Chin Med J (Engl) 116:1074–1077
Hall NM, Brown GM, Furlong RA et al (2003) Usp9y (ubiquitin-specific protease 9 gene on the Y) is associated with a functional promoter and encodes an intact open reading frame homologous to Usp9x that is under selective constraint. Mamm Genome 14:437–47
Faus H, Meyer HA, Huber M et al (2005) The ubiquitin-specific protease USP10 modulates androgen receptor function. Mol Cell Endocrinol 245:138–146
Bosch-Comas A, Lindsten K, Gonzalez-Duarte R et al (2006) The ubiquitin-specific protease USP25 interacts with three sarcomeric proteins. Cell Mol Life Sci 63:723–734
Valero R, Bayes M, Francisca Sanchez-Font M et al (2001) Characterization of alternatively spliced products and tissue-specific isoforms of USP28 and USP25. Genome Biol 2:RESEARCH0043
Xu J (2005) Age-related changes in Usp9x protein expression and DNA methylation in mouse brain. Brain Res Mol Brain Res 140:17–24
Dasari VK, Goharderakhshan RZ, Perinchery G et al (2001) Expression analysis of Y chromosome genes in human prostate cancer. J Urol 165:1335–1341
Soncini C, Berdo I, Draetta G (2001) Ras-GAP SH3 domain binding protein (G3BP) is a modulator of USP10, a novel human ubiquitin specific protease. Oncogene 20:3869–3879
Valero R, Marfany G, Gonzalez-Angulo O et al (1999) USP25, a novel gene encoding a deubiquitinating enzyme, is located in the gene-poor region 21q11.2. Genomics 62:395–405
Ohsumi Y (2001) Molecular dissection of autophagy: two ubiquitin-like systems. Nat Rev Mol Cell Biol 2:211–216
Zheng Z, Cai C, Omwancha J et al (2006) Sumo-3 enhances androgen receptor transcriptional activity through a sumoylation-independent mechanism in prostate cancer cells. J Biol Chem 281:4002–4012
Egerer T, Martinez-Gamboa L, Dankof A et al (2006) Tissue-specific up-regulation of the proteasome subunit beta5i (LMP7) in Sjogren’s syndrome. Arthritis Rheum 54:1501–8
Altun M, Galardy PJ, Shringarpure R et al (2005) Effects of PS-341 on the activity and composition of proteasomes in multiple myeloma cells. Cancer Res 65:7896–7901
Voorhees PM, Dees EC, O’Neil B et al (2003) The proteasome as a target for cancer therapy. Clin Cancer Res 9:6316–6325
Acknowledgements
We want to thank Dr. Hu Weimin and Prof. Zhengwei Yang for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Deng, S., Zhou, H., Xiong, R. et al. Over-expression of genes and proteins of ubiquitin specific peptidases (USPs) and proteasome subunits (PSs) in breast cancer tissue observed by the methods of RFDD-PCR and proteomics. Breast Cancer Res Treat 104, 21–30 (2007). https://doi.org/10.1007/s10549-006-9393-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-006-9393-7